This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at the University of Wisconsin-Madison.
What are domains?
A domain is a stretch of amino acids that form a distinct unit of tertiary structure which can exist separately from the rest of the protein. A domain is not to be confused with the concept of DNA motifs, which are short recurrent patterns that may be a part of domains, but are not associated with a particular function themselves.
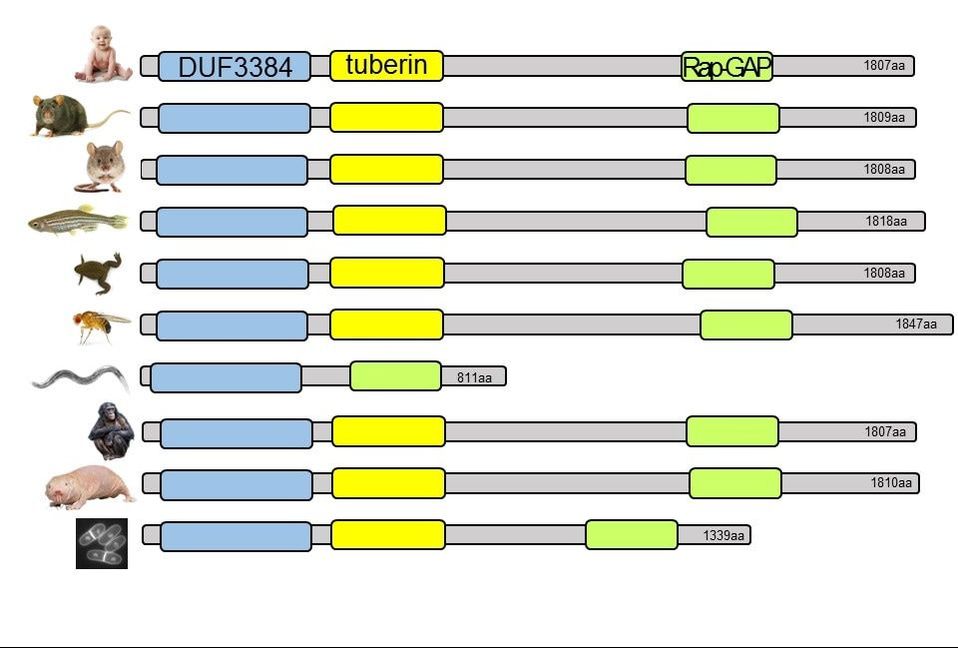
Protein domains are found through homology, particularly orthologs. These relationships are used to build protein families, which are organized based on similarity. Using the tools Pfam, SMART, InterPro, and PROSITE, domains were compiled for tuberin. Pfam, which looks for domains with a wide array of functions, was able to identify domains in both human tuberin and its rat homolog, thus it was used to compile results for the rest of the tuberin homologs. These domains included tuberin, Rap-GAP, and DUF3384. Below are the results as depicted by Pfam and SMART for human tuberin and its rat homolog.
|
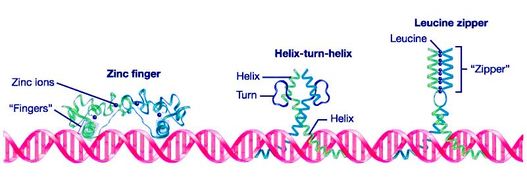
Tuberin Domains DUF3384: Includes a leucine zipper and coiled-coil motif. This is where tuberin binds to hamartin (TSC1) [1]. Tuberin: Includes a coiled-coil motif [1]. Rap-GAP: Tuberin has GTPase-activating protein activity toward the ras family small GTPase rheb, thus increasing hydrolysis of rheb and decreasing downstream signaling [1]. |
How well conserved is tuberin across species?
Discussion
|
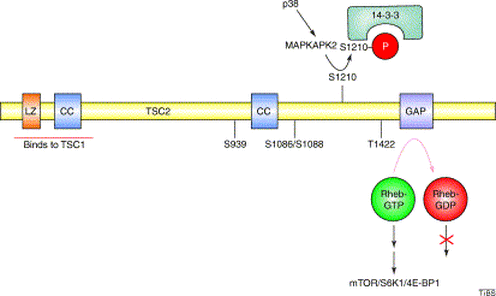
As seen above, tuberin protein domains are highly conserved among most of the selected organisms. Protein domains are important because they can predict important information about protein function with just its sequence. This is also useful when considering effects of mutations, where mutations in certain domains are likely to cause a corresponding loss of function. Additionally, knowledge of domains can be useful in experiments to isolate specific roles of protein. Below is a figure depicting the functions of tuberin's domains [1].
|
References
[1] Li, Yong, Corradetti M. N., Inoki, K., Guan, K. (2004). TSC2: filling the GAP in the mTOR signaling pathway. Trends in Biochemical Sciences 29(1), 32-38. http://dx.doi.org/10.1016/j.tibs.2003.11.007
Image References
Header: https://wallpaperscraft.com/image/smoke_paint_sheet_form_15699_1920x1080.jpg
Fig 1. Pfam, SMART
Fig 3. Li, Yong, Corradetti M. N., Inoki, K., Guan, K. (2004). TSC2: filling the GAP in the mTOR signaling pathway. Trends in Biochemical Sciences 29(1), 32-38. http://dx.doi.org/10.1016/j.tibs.2003.11.007
[1] Li, Yong, Corradetti M. N., Inoki, K., Guan, K. (2004). TSC2: filling the GAP in the mTOR signaling pathway. Trends in Biochemical Sciences 29(1), 32-38. http://dx.doi.org/10.1016/j.tibs.2003.11.007
Image References
Header: https://wallpaperscraft.com/image/smoke_paint_sheet_form_15699_1920x1080.jpg
Fig 1. Pfam, SMART
Fig 3. Li, Yong, Corradetti M. N., Inoki, K., Guan, K. (2004). TSC2: filling the GAP in the mTOR signaling pathway. Trends in Biochemical Sciences 29(1), 32-38. http://dx.doi.org/10.1016/j.tibs.2003.11.007