This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at the University of Wisconsin-Madison.
What is gene ontology?
The word ontology itself arose from the philosophical study of the nature of being and its categories. Therefore it makes sense that the Gene Ontology Consortium would adopt this as its name when it arose in an attempt to categorize and define relationships between various genes and gene functions to form a framework for biology. There are three parts to this, including the cellular component, molecular function and biological process.
Header: Bram van Velde, Hatching, 1980
Tuberin Ontology
Name: TSC1-TSC2 complex
Ontology: cellular_component
Synonyms: tuberin-hamartin complex, tuberous sclerosis complex
Definition: A protein complex consisting of at least tumerin and hamartin; its formation may regulate hamartin homomultimer formation. The complex acts as a GTPase activating protein (GAP) for the small GTPase (Rheb), and inhibits the TOR signaling pathway. Source: PMID:10585443, PMID:9580671, PMID:17121544
Name: TSC1-TSC2 complex
Ontology: cellular_component
Synonyms: tuberin-hamartin complex, tuberous sclerosis complex
Definition: A protein complex consisting of at least tumerin and hamartin; its formation may regulate hamartin homomultimer formation. The complex acts as a GTPase activating protein (GAP) for the small GTPase (Rheb), and inhibits the TOR signaling pathway. Source: PMID:10585443, PMID:9580671, PMID:17121544
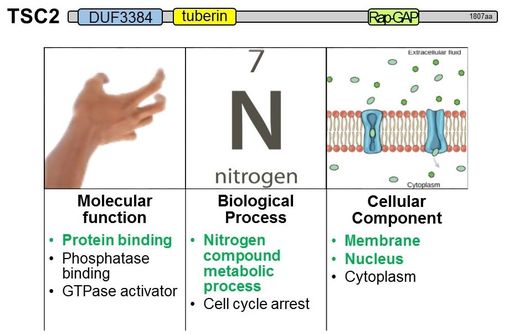
DiscussionAs seen above, a single protein often plays many roles and is involved in many different processes. Gene ontology is a useful tool for bringing the big picture together. Tuberin is involved in many processes including protein import into the nucleus, regulation of endocytosis, negative regulation of the insulin receptor signaling pathway, negative role of PI3K signaling, cell cycle arrest, neural tube development and closure, heart development, and many more. This makes the multitude of characterizations of tuberous sclerosis complex seem very reasonable.
Additionally, gene ontology can be useful because it brings together the multitude of discoveries about single proteins together in one place. By doing so, relatively unexplored areas can be identified. For example, it was found that tuberin is associated with nitrogen metabolism, but the role of tuberin in nitrogen metabolism in the kidney is currently unknown. |
References
Image References
Header: http://theredlist.com/wiki-2-351-861-414-1293-1237-1291-view-european-abstraction-profile-van-velde-bram.html
Fig 1: http://www.cse.buffalo.edu/~rapaport/663/F08/krresources.html
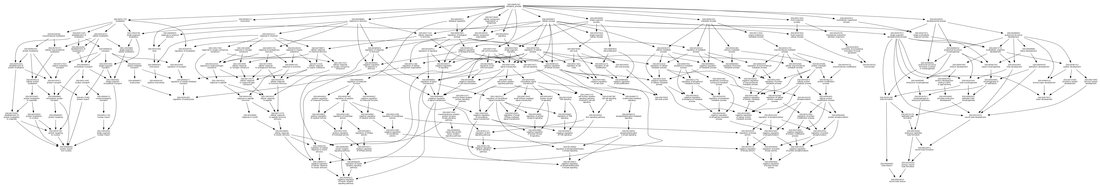
Fig 2: http://biit.cs.ut.ee/gprofiler/index.cgi?organism=hsapiens&sort_by_structure=1&query=TSC2&significant=0&output=png
Image References
Header: http://theredlist.com/wiki-2-351-861-414-1293-1237-1291-view-european-abstraction-profile-van-velde-bram.html
Fig 1: http://www.cse.buffalo.edu/~rapaport/663/F08/krresources.html
Fig 2: http://biit.cs.ut.ee/gprofiler/index.cgi?organism=hsapiens&sort_by_structure=1&query=TSC2&significant=0&output=png